
The table and heatmap views are fully customizable by selecting the Customize View option. Any missing data points will be a yellow color. This shows a Heatmap view of the gene abundance in different samples.
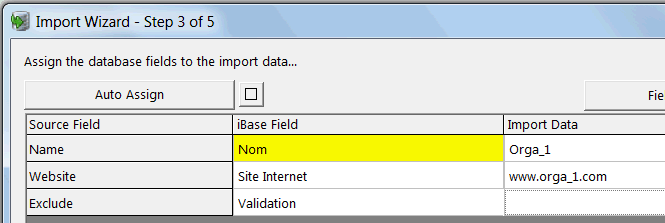
Individual genes can also be deleted by first clicking on the gene ID (header of the row) and clicking on Remove selected assays.Ĭlick the HeatmapTable tab to see the heatmap view of the data. Notice that number of sample selected will be updated to reflect how many you have selected. Samples can be removed in this screen by clicking on sample IDs on the column header and clicking on Remove selected samples option. The right side frame contains statistics for the data being imported, as well as visualizations (Heatmap Table tab) of the data. Workflow frame of the Import RT-PCR Wizard window now shows that we have advanced to the Preview data step. Select the Next button at the bottom of the window Preview Data
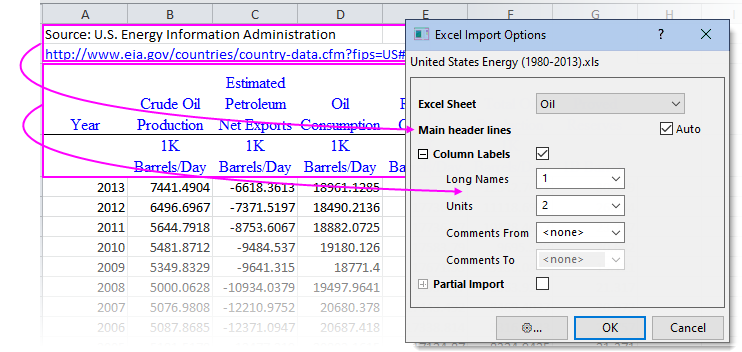
Uses can also import TaqMan hPSC Scorecard data, formatted as below: Uses can also import QuantStudio RT-PCR data.Īs one row is for one data point, the data can be imported as Tall Skinny Data format. The user also has the option to import the original CT data into the project. The only option that could possibly be in need of changing at this stage is the Remove assays with no data points and Remove samples with no data points options. Table data (one gene per row) - For this format, data should be in a table where each row contains the information for all the genes and may include other descriptive information for the assay.īecause the data was imported using the Manage RT-PCR Raw Data module, most of the options on this screen will be filled in by detecting the file automatically. Table data (one sample per row)- For this format, data should be in a table where each row contains the information for all the assays and may include other descriptive information for the sample. Tall skinny data (one data point per row)- For this format, data should be in a table where each row contains the information for a well, and may also include other descriptive information for the sample and assay. This is the format used in Tools | Data | Manage Taqman Raw Data. This table should include the following columns #, PlateID, Well, AssayID, SampleID, Value, and Include. Standard format (one data point per row)-For this format, Data should be a table containing exactly 7 columns, including a row header column. Also, clicking the Workflow button will allow the user to save the workflow to return at another time.Īrray Studio supports importing data from the following formats: Initially, this will be the Import data step, as shown below but will be updated throughout the completion of the wizard. The Workflow section shows the user on which step of the Import RT-PCR Wizard the user is working. The Import RT-PCR Wizard is now displayed and contains two main sections. In this tutorial, since the imported plate file did not contain abundances, choose the first option and click OK now: Using the Ct/Cp values, using the abundance values, or using any available abundance values and converting Ct/Cp to abundance where abundance is not available. It can be opened via the Add RT-PCR Data menu item, or through the Manage RT-PCR Raw Data menu (after adding plate files).Ĭlick the File | Import RT-PCR Wizard to continue:Īt this point, the user is asked to choose an RT-PCR Value source. The Import RT-PCR Wizard can be used to walk the user through the process of importing and normalizing data. Marker Statistics Data Filtering and Population StructureĬonnecting To A Server And Uploading Files Connecting to a Server and Uploading Files
